Cells spend most of their time doing two things: growing or not growing. This is so obvious that it’s easy to forget the profound differences between these states. Growing cells have an intense demand for nutrients and energy necessary to drive biosynthesis. Quiescent cells need only maintain stasis. Most remarkably, cells can quickly flip between these states when growth signals fluctuate, triggering a system-wide reprogramming of nearly all aspects of cellular metabolism. The precise execution of this “growth program” is a fundamental determinant of the sizes of cells (and tissues and organisms), how fast they grow, and how energy is consumed and used. Loss of growth control is a common cause of human disease, especially in cancers and metabolic disorders. Our lab is seeking to understand the molecular mechanisms that orchestrate this program, their function in normal growth, and how dysfunction drives human disease. We are particularly interested in post-transcriptional control of gene expression by mechanisms that modulate mRNA translation and stability.
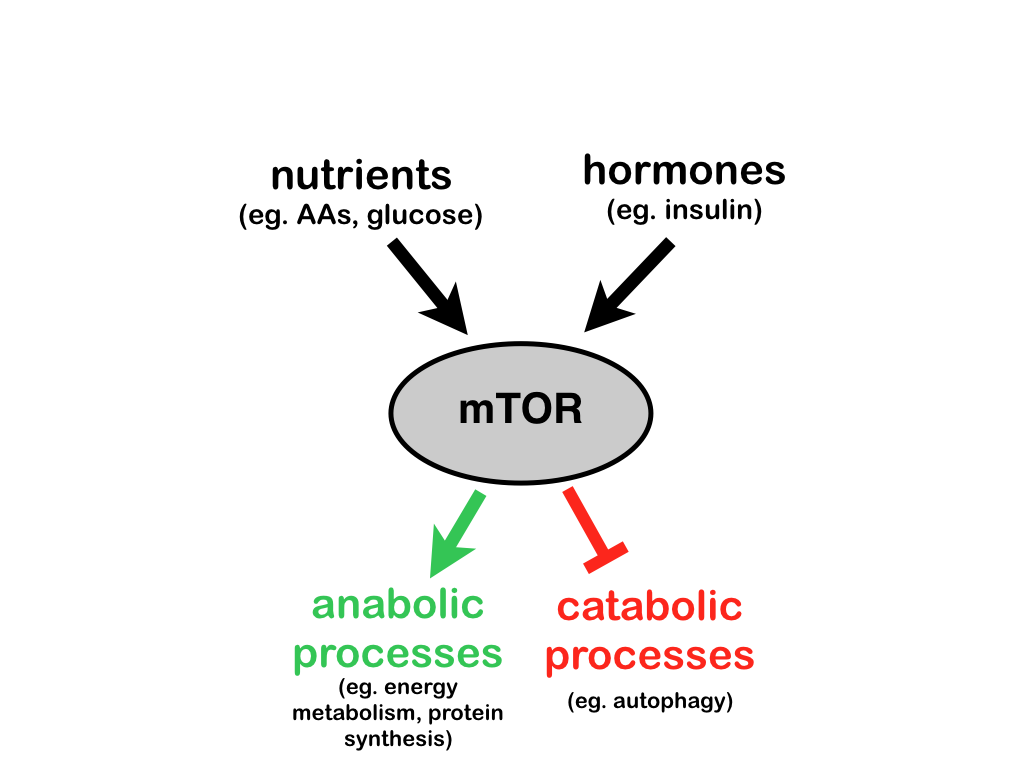
A major focus of our current research is the mTOR signaling pathway, a sensor of the cellular nutrient status and a master regulator of cell growth. This pathway is a major regulator of mRNA translation, and is deregulated is a wide variety of diseases, including cancer, metabolic disease and neurologic disorders. See a review here. The following projects are ongoing:
Translational control by the mTOR pathway
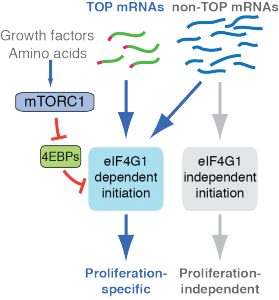
We previously found that acute inhibition of mTOR selectively inhibits the translation of large class of mRNAs containing a terminal oligopyrimidine (TOP) motif at the 5’ terminus by disrupting the mRNA cap-binding complex, eIF4F. The mTOR pathway has many additional targets in the translational machinery, but the functional significance of these is unknown. We want to understand how mTOR-regulated translational mechanisms work in molecular detail, what features in mRNAs determine their dependence on mTOR activity, and how these controls are employed physiologically.
Growth-dependent control of mRNA stability
The switch from growth to quiescence changes mRNA levels throughout the transcriptome. These changes are precipitated by regulation of both mRNA production and decay. We are developing new ways to measure these parameters throughout the transcriptome and determine how mTOR controls them.