Have you ever wondered how cells can change shape so rapidly and so robustly? How molecular assemblies, such as the clathrin-mediated endocytosis machinery, can deform the plasma membrane so dramatically that they transform a flat surface into a vesicle in less than 10 seconds?
If you’re not already convinced these are not an easy problems, try this quick experiment. Take a piece of fabric, let’s say the sleeve of your hoodie. Now, with one hand, try to deform it to make a small vesicle, or just a tubule…
Not easy huh? Eukaryotic cells, like yeast, have figured out how to solve this problem more than a billion years ago! “Just” assemble the right copy number of the right molecules, at the right place, at the right time…
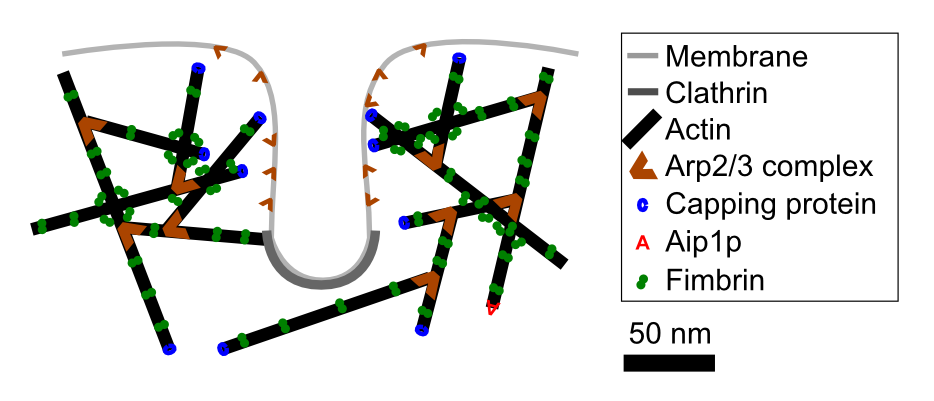 The research in the Berro lab aims to understand the dynamics of these molecular machineries, how they generate forces to deform membranes and how they adapt to different biophysical and biochemical conditions and make the process robust. We are particularly interested in the dynamics of clathrin-mediated endocytosis.
The research in the Berro lab aims to understand the dynamics of these molecular machineries, how they generate forces to deform membranes and how they adapt to different biophysical and biochemical conditions and make the process robust. We are particularly interested in the dynamics of clathrin-mediated endocytosis.
To address these questions, we use a combination of experimental, computational and theoretical approaches. Our research relies on live quantitative microscopy, a method to count the copy number of proteins in specific location in live cells. Combined with advanced data analysis and mathematical modeling, such temporal datasets are an invaluable source of information for inferring the function of individual proteins in molecular assemblies.
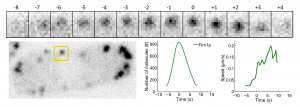
Quantitative microscopy of live cells allows us to follow the temporal evolution of the absolute number of molecules in individual endocytic patches, along with its speed. Top row: temporal evolution of an individual endocytic event (orange box in the cell below). Bottom row: temporal evolution of the copy number of fimbrin and of the speed of endocytic vesicles
Our model system is fission yeast, Schizzosaccharomyces pombe, because this haploid eukaryot has the same core molecular machinery for clathrin-mediated endocytosis as mammals. Tools for yeast genetics are extremely efficient and fast. In addition, the fission yeast genome is much smaller (about 5000 genes) than mammalian genomes, which make it much more tractable. Working with with yeast is so easy, even a mathematician can do it!


