 https://orcid.org/0000-0002-0816-6115
https://orcid.org/0000-0002-0816-6115
Lake Baikal, Epischurella (Epischura) baikalensis, and climate change
I have been engaged in research in the Lake Baikal system for just over 7 years. The future of Lake Baikal’s biodiversity is uncertain with climate rapidly changing in the region. Unlike its diverse benthos, Lake Baikal’s zooplankton is species-poor, with up to 90% of its biomass being composed of a single Calanoid copepod species, Epischurella baikalensis, renamed from Epischura baikalensis here.

My research has characterized the genetic differentiation and differential gene expression of E. baikalensis across the lake’s massive expanse. Using pooled transcriptomics, we identified genetic differentiation and differential gene expression in many genes, including those related to biochemical pathways involved in heat shock, found here.

Dimensions of Biodiversity: Lake Baikal Research Blog
Daphnia Population Dynamics and Genetics
A significant portion of my dissertation research investigated the evolution of life history traits, particularly in response to changes in temperature. As climate change increase in inland waters, especially in the temperate region, it is important to know how those changes will trickle through ecosystems. I used lakes and kettle ponds in coastal Connecticut to
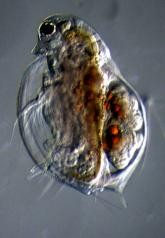
understand how changes in individual zooplankton metabolism affects the population, community, and ecosystem. I worked in the lab with Daphnia ambigua, D. pulicaria, and D. dentifora and applied those outcomes to field systems. In the field, I used Bride, Dodge, and Gorton ponds as naturally occurring replicates to understand how individual metabolism and ecosystem metabolism are linked. Whereas, I use controlled laboratory growth chambers to understand the specific effects of climate on zooplankton physiology. To do this, I use RNA:DNA ratios, a method pioneered in the 1980s (that I revisited during during my master’s thesis), which was used to assess fish population starvation. My goal is to be able to connect those concepts into a deeper understanding of what to expect if natural aquatic ecosystems continue to warm rapidly.
I also completed a global meta-analysis of Daphnia life history traits to understand the contributions of the environment, temperature, and evolutionary history on the evolution of fitness related traits. The results of these projects are forthcoming.
STEM Education
While at ETSU, I was the recipient of the National Science Foundation’s Graduate K-12 Fellowship. Under this fellowship, I was challenged with bringing my genetics research to local pre-kindergarteners and kindergarteners.
As I became well-versed in the Tennessee State Science curriculum, I became interested in how Tennessee (and other states in the Southeast) would adapt and change to incorporate the new Next Generation Science Standards (2011), the suite of Science curriculum standards released by the National Research Council in tandem with the Common Core State Standards for Mathematics and English and Language Arts.
I spent a significant amount of time understanding how science is taught in the K-12 setting and how the NGSS would allow science curricula across the country to grow and develop over time.
Understanding best practices in relaying information from primary sources down to the K-12 setting is and will always remain an important motivation for my outreach and research. See more here, here, and here.
I also contributed to an amazing team developing tools for prefaculty teaching training, where we were able to show that even small course-based bootcamp interventions make demonstrable differences to how graduate students and postdocs make their classrooms inclusive and feel about their own teaching abilities. Results forthcoming.

